|
The WES data was derived from each tumor to a median of 260x depth. We also performed RNA sequencing (RNA-seq) for the 33 ESCC patients with high RNA quality. We calculated CpG-rich methylation on a genome-wide scale for all the patients by single-cytosine-resolution DNA methylation analysis using reduced representation bisulfite sequencing (RRBS) (Fig. All samples (including adjacent normal tissues) were characterized by whole-exome sequencing (WES). In this study, we prospectively collected 186 samples from 36 ESCC tumors from 36 patients (Supplementary Fig. Indeed, although a strong correlation between TCR (T cell receptor) repertoire and genomic ITH has been found in ESCC, mechanisms of neoantigen evasion in ESCC remain unclear. Moreover, the interaction between the cancer cell and its immune microenvironment and how their cross-talk influences cancer evolution has not been investigated. Thus, an understanding of the complex interplay between the genome, transcriptome, and methylome underpinning ITH in ESCC and the evolution of this disease is lacking. However, ESCC ITH has not been comprehensively characterized across multi-omics from a large cohort of ESCC patients. Recently, several studies have performed multi-regional whole-exome sequencing (WES), methylation profiling, or TCR sequencing to unveil ITH in ESCC 10, 11, 12. A precise understanding of ITH is crucial for the development of effective diagnosis and biomarker design. ITH, which supplies the fuel for clonal evolution and drug resistance 9, is pervasive across multiple levels of molecular features. However, oncological biomarkers might be confounded by sampling bias derived from spatial intratumoral heterogeneity (ITH). 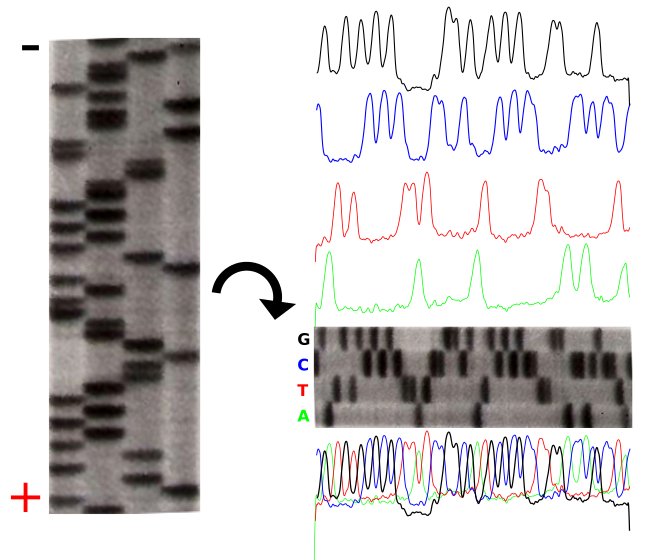
Several large-scale sequencing studies have revealed the complex genomic landscape of ESCC 3, 4, 5, 6, 7, 8. Overall, our work offers an avenue of dissecting the complex contribution of the multi-omics level to the ITH in ESCC and thereby enhances the development of clinical therapy.Įsophageal squamous cell carcinoma (ESCC) is one of the most prevalent cancer types which occurs in Eastern Asia and parts of Africa 1, 2. We reveal genetic mechanisms of neoantigen evasion under distinct selection pressure from the diverse immune microenvironment. Based on the RNA-seq data, we observe diverse levels of immune infiltrate across different tumor sites from the same tumor. Through multi-omics analyses, it is discovered that genomic, epigenomic, and transcriptomic ITH are underpinned by ongoing chromosomal instability. In this study, we perform multi-omic sequencing on 186 samples from 36 primary ESCC patients. Further, the extent to which tumor-infiltrated T lymphocytes are directed against cancer cells, but how the immune infiltration acts as a selective force to shape the clonal evolution of ESCC is unclear. However, although genomic ITH has been characterized in genetic, transcriptomic and epigenetic alterations are hallmarks of esophageal squamous cell carcinoma (ESCC), the extent to which these are heterogeneous in ESCC has not been explored in a unified framework. Meaning that adding steps won't stop the process.Īctually, they work on different principles and do not know how to exactly answer your question.Intratumoral heterogeneity (ITH) has been linked to decreased efficacy of clinical treatments. This allows you to 'physically' separate the signal originating from each target molecule. What is really important to understand is that each cluster is generated from a SINGLE MOLECULE of target DNA. 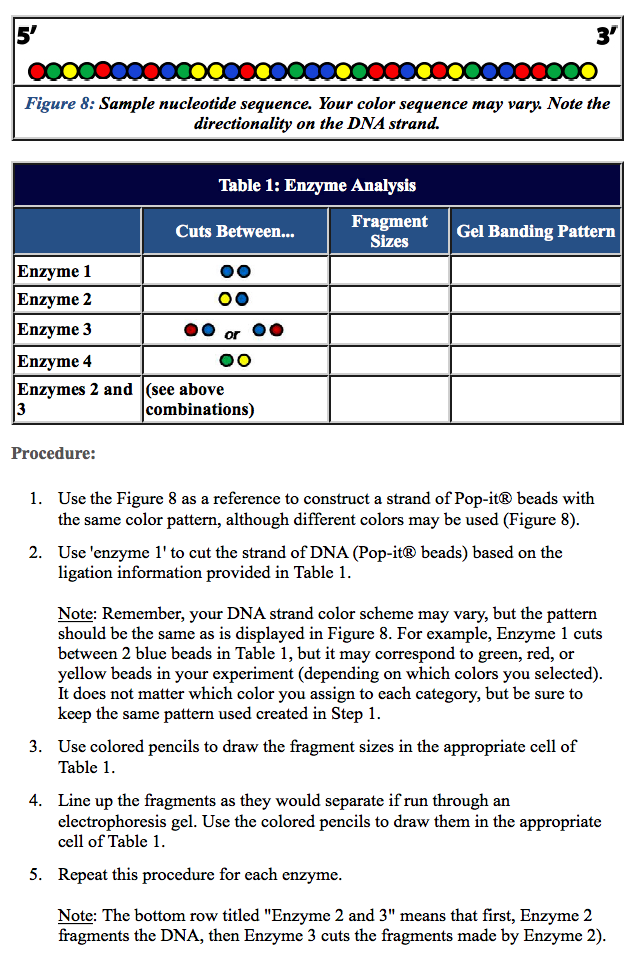
With NGS (Illumina in this example but the concept is the same for other NGS technologies) you prepare a target-DNA library and you load it into a flowcell in order to generate DNA clusters that are then sequenced in parallel. In Sanger Sequencing, dideoxynucleotides are added to stop sequencing and help us read the products. The difference between next-generation sequencing and Sanger sequencing is that NGS allows us massive parallel sequencing. They all rely on adding nucleotides to continue the reaction.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
